 |
By Christina Towers, PhD
A novel vesicular organelle was recently discovered in 2015 by a research group at Tsinghua University headed by Dr. Yu that is dependent on cell migration and named accordingly as migrasomes1. They are classified as extracellular vesicles formed on the intersections of retraction fibers trailing behind migrating cells. High resolution imaging showed that tiny vesicles combine to make larger structures that grow up to 3μM in diameter and eventually break off the fibers into the extracellular space. Isolation with density-gradient centrifugation and subsequent mass spectrometry revealed an abundance of membrane-localized and cytoskeleton-related proteins, and especially high levels of tetraspanin 4. Accordingly, confocal imaging has been used to visualize fluorescently tagged tetraspanin 4 enriched on migrasomes. The formation of migrasomes is dependent on cell migration. Genetic and/or pharmacological manipulations that increase cell migration caused a corresponding increase in the number of migrasomes, while inhibition of cell migration decreased migrasome formation. Perhaps most intriguing is that migrasomes can be taken up by surrounding cells, suggesting they may play a role in cell to cell communication.
While this first publication showed migrasomes exist in multiple cell types including human cancer cell lines, cultured neurons, and macrophages, it was unclear what, if any, the function of these novel vesicles might be in vivo. In a new publication released by the same group in 2019, Jiang et al utilized zebrafish as a model organism to investigate the role of migrasomes during embryogenesis – a process dependent upon complex cell migration patterns2. Confocal imaging of zebrafish embryos undergoing gastrulation, at approximately 6 hours post fertilization (hpf), which were injected with GFP labeled tetraspanin 4 mRNA showed numerous migrasomes forming on refractory fibers. Further temporal-spatial analysis showed the number of migrasomes increased dramatically between 5.5-7 hpf, and most migrasomes were localized to pockets of extracellular space between cells. Embryos with CRISPR/Cas9 mediated knockout of tetraspanin 4 or tetraspanin 7, tspan4a−/− or tspan7−/−, formed fewer migrasomes during gastrulation. Interestingly, despite normal gastrulation in these mutants, at 48hpf 65% of the embryos presented impaired organ morphogenesis. Moreover, mass spectrometry analysis from migrasomes isolated from over 20,000 embryos showed a more than two-fold enrichment in many chemokines, morphogens and growth factors compared to cell bodies, further suggesting a signaling role for these organelles.
|
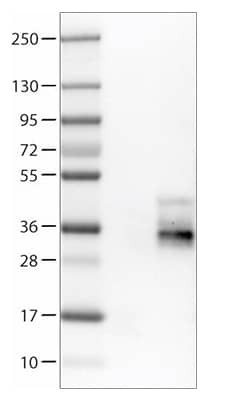 |
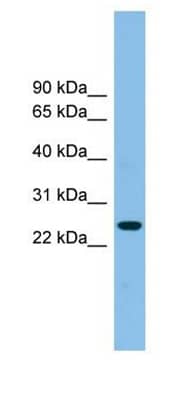
Western blot analysis of (Left) Tetraspanin-4 in Jurkat cell lysate using rabbit polyclonal tspan4 antibody [NBP1-59438]. |
|
With only a handful of publications describing these novel vesicles, there are still a lot of open questions. These data suggest a role in cell to cell signaling, but the details of those mechanisms are yet to be worked out. Given that most cell types make migrasomes, it is likely they may play a role in disease pathology, particularly in processes dependent on cell migration. For example, one might hypothesize a casual role for migrasomes in cancer cell migration, invasion, and eventual metastasis. Nonetheless, the discovery of these novel vesicles opens the door for many future studies to fully elucidate their mechanism of formation, roles during development, and function in disease pathology.
 Christina Towers, PhD
Christina Towers, PhD
University of Colorado (AMC)
Dr. Towers studies the roles of autophagy, apoptosis and cell death in cancer.